GeneStrands Fragments perform better, are delivered faster, and cost the same as or lower than gene fragments from other suppliers.
GeneStrands are linear, double-stranded DNA fragments constructed using synthetic DNA and are ready for use in applications such as cloning, CRISPR-based genome editing, and antibody engineering. GeneStrands serve as an inexpensive substitute for our gene synthesis service.
- Sequences up to 3,000 bp
- Delivered in liquid form as a dsDNA fragment (min. 200 ng)
Order a GeneStrand with desired embedded restriction sites and construct your gene of interest in your own lab by cloning directly into your vector of choice using TA cloning, Gibson assembly, or other gene construction protocols.
Pricing and Turnaround Time
| Length |
Tube Price |
Delivery
(business days) |
Plate Price |
Delivery
(business days) |
Amount (ng) |
| 100–500 bp |
$100.00 USD |
2–4 |
$90.00 USD |
8-10 days |
300 |
| 501–750 bp |
$144.00 USD |
2–4 |
$100.00 USD |
8-10 days |
300 |
| 751–1000 bp |
$170.00 USD |
3–5 |
$112.00 USD |
8-10 days |
300 |
| 1001–1250 bp |
$237.00 USD |
5–8 |
$180.00 USD |
8-10 days |
300 |
| 1251–1500 bp |
$283.00 USD |
5–8 |
$226.00 USD |
8-10 days |
300 |
| 1501–1750 bp |
$330.00 USD |
5–8 |
$258.00 USD |
8-10 days |
300 |
| 1751–2000 bp |
$397.00 USD |
5–8 |
$314.00 USD |
8-10 days |
300 |
| *2001-2250 bp |
$474.00 USD |
5–8 |
$386.00 USD |
8-10 days |
300 |
| *2251-2500 bp |
$530.00 USD |
5–8 |
$427.00 USD |
8-10 days |
300 |
| *2501-2750 bp |
$597.00 USD |
5–8 |
$484.00 USD |
8-10 days |
300 |
| *2751-3000 bp |
$654.00 USD |
5–8 |
$530.00 USD |
8-10 days |
300 |
View all pricing and turnaround for genes.
Order Now
Why Eurofins?
- The fastest turnaround times in the industry.
- Since GeneStrands are assembled using synthetic DNA oligos, the sequences of GeneStrands gene fragments, much like the sequence of our regular genes, can be optimized for expression in your host of choice.
- The free in-silico sequence optimization tool GENEius, which is integrated with the order wizard, can optimize the sequence to remove complex segments, remove/introduce specific restriction sites, and optimize the sequence for heterologous expression.
- Assembled using oligos that are synthesized using high-fidelity chemical synthesis protocol and ensure high Cloning Efficiency and Sequence Accuracy in the clones.*
- Confidentiality of your sequence.
- Discounts are available for high volume orders or with long term contracts. Please contact a sales representative for further details.
*use of error-removal methods during assembly (which we rountinely employ for the construction of GeneStrands and Genes) reduces this error significantly.
Molecular Cloning with GeneStrands
All Gene Fragments are synthesized under the same meticulous conditions as our gene synthesis service. However, Gene Fragments are delivered as uncloned products, and as such the presence of minute populations of truncated or mutated species is unavoidable. We recommend that after cloning a GeneStrand with a regular sequence, select multiple individual colonies and verifying sequence after each cloning step. For GeneStrands with complex segments in the sequence, we recommend selecting a higher number of colonies for sequence verification. Our scientists select <10 colonies for mini-prep; typically, 4–6 of them contain the vector with right-sized insert and the desired error free sequence.
Assembly of GeneStrands gene fragments at Eurofins Genomics.
Step 1:
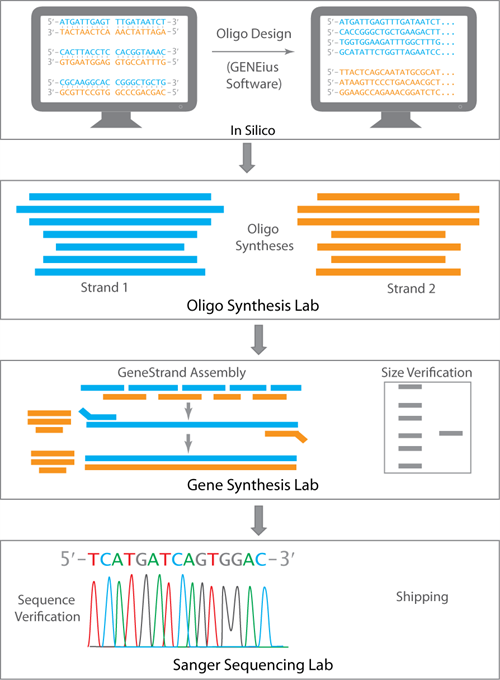 Once the order has been submitted the propreitary GENEius software tool breaks the sequence down to smaller oligos to ensure that there is sufficient overlap between contiguous oligos and the complementary bridging splint.
Once the order has been submitted the propreitary GENEius software tool breaks the sequence down to smaller oligos to ensure that there is sufficient overlap between contiguous oligos and the complementary bridging splint.
Step 2:
The designed oligos are then synthesized at our DNA Synthesis Lab using high-fidelity chemical synthesis - very high coupling efficiency and mild post-synthesis clean up. The mass spec. validated oligos are then used in the assembly of the dsDNA.
Step 3:
The oligos are processed, annealed, and then ligated to contruct one strand of the full length DNA. High-Fidelity DNA polymerases are then used to generate the second strand and amplify the dsDNA. The size of the resulting PCR product is verified by gel electrophoresis using a gel with the appropriate resolution.
Step 4:
The size-verified DNA is then dried down and shipped as GeneStrand to you. The size-verified dsDNA can be sequenced at our sequencing lab for additional confirmation. Sequence validation is of the entire pool, so the results obtained are a representation of the predominant sequence within the pool. Sequence information is kept confidential at our facility. A copy of the sequence will then be available for a $25 fee.
We ensure strict confidentiality of every project. The high quality standard of our service is assured by a professional quality management team. The quality management system of our Louisville, Kentucky facility is certified to ISO 9001:2008 and ISO 13485:2003 quality standards.
Sequence optimization (optional)
Sequences of high complexity may prolong the overall turnaround time of delivery and, in some extreme cases, disqualify the sequence for delivery as GeneStrand. Please note that we offer free in silico sequence optimization option on our website. This optimization can eliminate sequence complexities, while increasing the yield the protein expressed in the host.
Complexities in a sequence
Complex segments in the sequence make synthesis of oligos and the assembly of GeneStrand gene fragments challenging, while also reducing the efficiency of transcription and translation. Complexities in a sequence include:
- Stretches of direct or inverted repeats (>20 bp)
- homopolymeric stretches (>18 bp)
- Stretches of sequence that can result in critical secondary structures
- Stretches of sequence with very high (>75%) or low (<35%) GC content.
Secondary structures reduce the synthetic yields of DNA oligos while also reducing the translational efficiency, since these secondary structures are then adopted by the mRNA. Any Genes or GeneStrands that contain these features are considered 'complex' and take longer to assemble.
85% cloning efficiency
Eurofins Genomics routinely tests the mutation rate and quality of its synthesized GeneStrands after inserting into a vector. We have observed that over 85% of the clones contain a DNA of the expected size when GeneStrands are used in thier assembly. This features ensures that, with GeneStrands, you have to select fewer colonies to obtain the correct gene of interest.
Quality Assurance
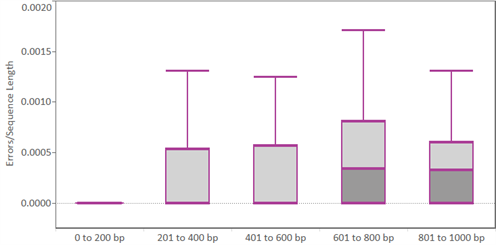 |
| Errors in GeneStrands: Box-and-whisker plots depicting the distribution of sequencing errors observed in GeneStrands after inserting them into vectors. 873 distinct GeneStrands of different lengths (>200 datapoints for sizes 0 to 200 bp, 201 to 400 bp, and 401 to 600 bp; and >50 datapoints for 601 to 800 bp and 801 to 1000 bp) from a period in 2014 were evaluated. No error-removal methods were used in the assembly of these GeneStrands. |
Order Now